A Lineweaver-Burk plot is a graphical representation of the Michaelis-Menten enzyme kinetics equation, which describes the relationship between the reaction rate(V) and the substrate concentration[S]. In this post, I will show you how to create a Lineweaver-Burk plot from Michaelis-Menten data with two examples.
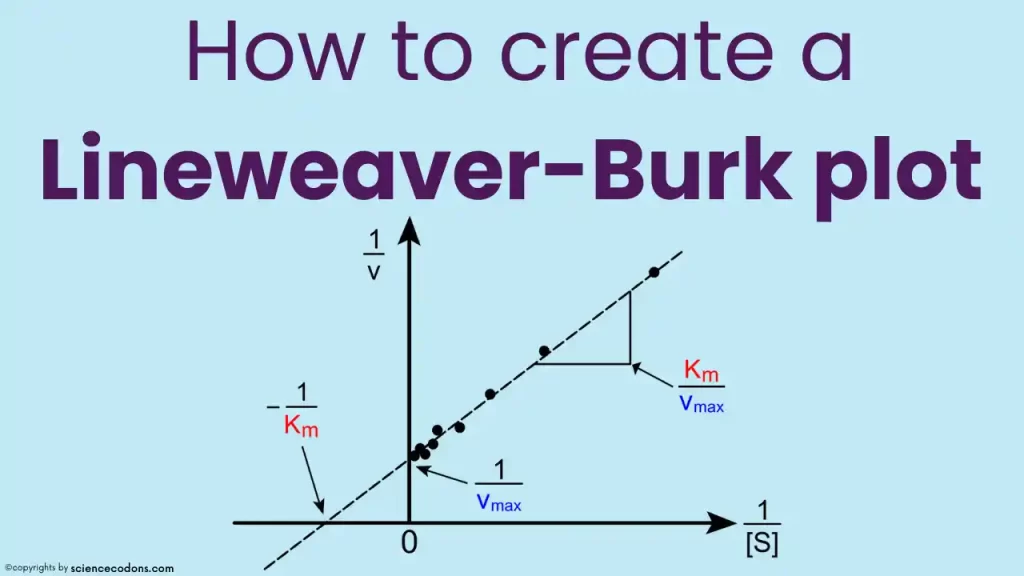
What is a Lineweaver-Burk plot?
The Michaelis-Menten equation is given by:
$$v = \frac{V_{max}[S]}{K_m + [S]}$$
Where \(v\) is the reaction rate, \(V_{max}\) is the maximum reaction rate, \(K_m\) is the Michaelis constant, and [S] is the substrate concentration.
To create a Lineweaver-Burk plot, we take the reciprocal of both sides of the equation and rearrange it as follows:
$$\frac{1}{v} = \frac{K_m}{V_{max}}\frac{1}{[S]} + \frac{1}{V_{max}}$$
This equation has the form of a straight line: \(y = mx + b\), where \(y = \frac{1}{v}\), \(x = \frac{1}{[S]}\), \(m = \frac{K_m}{V_{max}}\), and \(b = \frac{1}{V_{max}}\).
Therefore, by plotting \(\frac{1}{v}\) against \(\frac{1}{[S]}\), we can obtain a straight line whose slope and intercept can be used to calculate the values of \(K_m\) and \(V_{max}\).
How to create a Lineweaver-Burk plot
To create a Lineweaver-Burk plot, we need to have some experimental data on the reaction rate \(v\) and the substrate concentration [S] for a given enzyme. To draw Lineweaver Burk plot from raw data, we need to perform the following steps:
- Calculate the reciprocal of the reaction rate \(\frac{1}{v}\) and the substrate concentration \(\frac{1}{[S]}\) for each data point.
- Prepare a table with four columns: substrate concentration [S], \(v\), \(\frac{1}{[S]}\), and \(\frac{1}{v}\).
- Plot the values of \(\frac{1}{v}\) on the y-axis and \(\frac{1}{[S]}\) on the x-axis using a scatter plot.
- Draw a best-fit line through the data points using a linear regression method.
- Find the slope and the intercept of the line using the equation of the line or the regression output.
- Calculate the values of \(K_m\) and \(V_{max}\) using the formulas: \(K_m = \frac{V_{max}}{m}\) and \(V_{max} = \frac{1}{b}\).
Example 1
Suppose we have the following data of the reaction rate \(v\) and the substrate concentration (\([S]\) for an enzyme that catalyzes the conversion of A to B:
| \([S]\) (mM) | \(v\) (µM/min) |
|---|---|
| 0.1 | 2.5 |
| 0.2 | 4.2 |
| 0.4 | 6.7 |
| 0.8 | 9.1 |
| 1.6 | 11.8 |
| 3.2 | 14.2 |
| 6.4 | 15.6 |
To create a Lineweaver-Burk plot, we first calculate the reciprocal of the reaction rate \(\frac{1}{v}\) and the substrate concentration \(\frac{1}{[S]}\) for each data point:
| \([S]\) (mM) | \(v\) (µM/min) | \(\frac{1}{[S]}\) (1/mM) | \(\frac{1}{v}\) (min/µM) |
|---|---|---|---|
| 0.1 | 2.5 | 10 | 0.4 |
| 0.2 | 4.2 | 5 | 0.238 |
| 0.4 | 6.7 | 2.5 | 0.149 |
| 0.8 | 9.1 | 1.25 | 0.11 |
| 1.6 | 11.8 | 0.625 | 0.085 |
| 3.2 | 14.2 | 0.3125 | 0.07 |
| 6.4 | 15.6 | 0.15625 | 0.064 |
Then, we plot the values of \(\frac{1}{v}\) on the y-axis and \(\frac{1}{[S]}\) on the x-axis using a scatter plot:
![Lineweaver-Burk plot example 1]
As expected, we can see that the data points form a straight line. We can use a linear regression method, such as the least squares method, to draw the best-fit line. The equation of the line is given by:
$$\frac{1}{v} = 0.064 \frac{1}{[S]} + 0.062$$
The slope of the line is \(m\) = 0.064, and the intercept is \(b\) = 0.062. Using these values, we can calculate the values of \(K_m\) and \(V_{max}\) as follows:
$$K_m = \frac{V_{max}}{m} = \frac{1/b}{m} = \frac{1/0.062}{0.064} = 0.97 \text{ mM}$$
$$V_{max} = \frac{1}{b} = \frac{1}{0.062} = 16.13 \text{ µM/min}$$
Therefore, the Michaelis constant \(K_m\) for this enzyme is 0.97 mM, and the maximum reaction rate \(V_{max}\) is 16.13 µM/min.
Reference:
Lineweaver–Burk plot – Wikipedia